Cardiovascular Epigenomics
Hypertension is the leading cause of cardiovascular disease and premature death worldwide. Globally, over 1 billion people are diagnosed with hypertension and this number has doubled since 1990. Lowering blood pressure can cut the number of strokes by 35%–40%, heart attacks by 20%–25%, and heart failure by 50%, yet only a minority of those with hypertension are optimally managed.
There are a large number of anti-hypertensive therapies available, yet individual responses vary and efficacy remains a concern. The initiating causes and pathogenic mechanisms for hypertension are largely unknown and markers for prognosis of adult hypertension could improve prevention, diagnosis and disease management.
Investigating the interaction between genetic, epigenetic and environmental factors which play a role in the mechanisms underpinning pharmacogenomic therapeutic options for treatment of hypertension, our multidisciplinary research brings together expertise in genomics with collaborators in regenerative medicine (University of Galway) and nutrition (NICHE, Ulster University). We use a range of molecular biology, cellular assays, human studies and bioinformatic analysis across four broad themes:
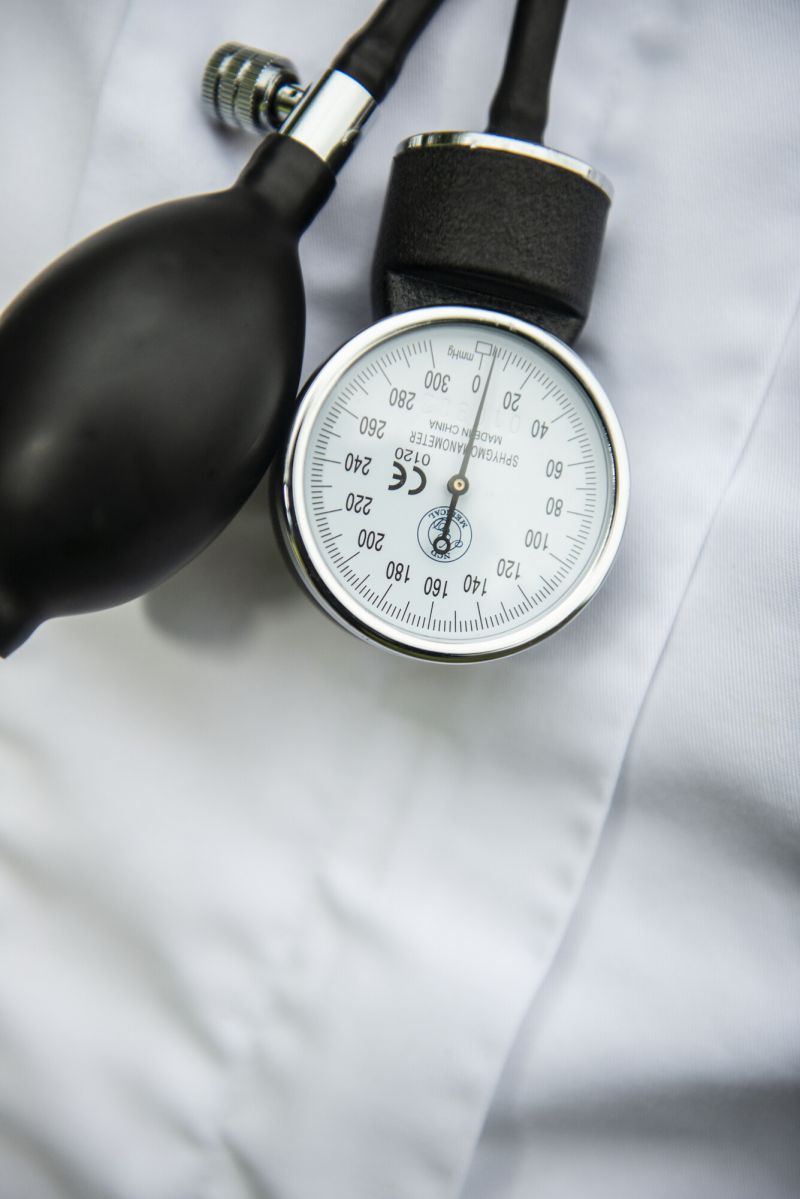
Epigenomics of Hypertension
Ongoing collaborative work with NICHE is providing novel insights into personalised hypertension management and treatment....

Regenerative approach to nutrigenomics of hypertension
Regenerative approaches have been used to successfully create patient-specific induced pluripotent stem cell (iPSC) models....
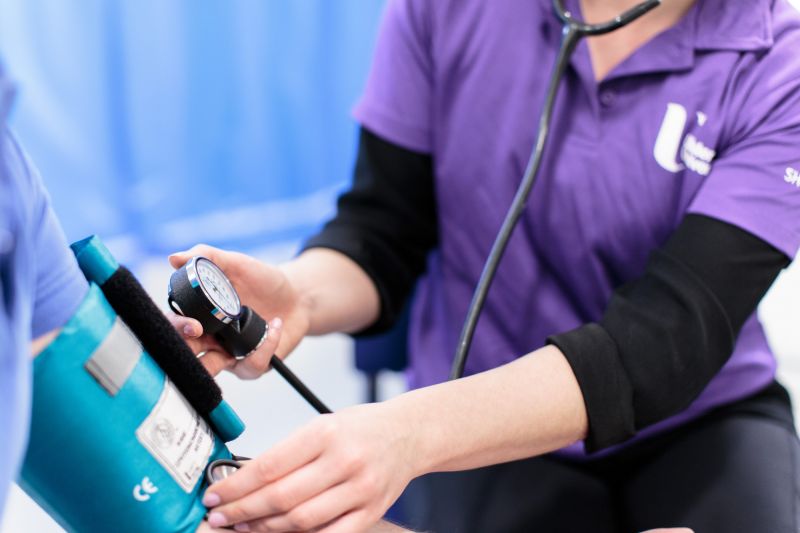
Multimodal data integration for identifying novel drug targets for hypertension
Hypertension (high blood pressure) effects one in three adults and is the number one cause of stroke in Northern Ireland....
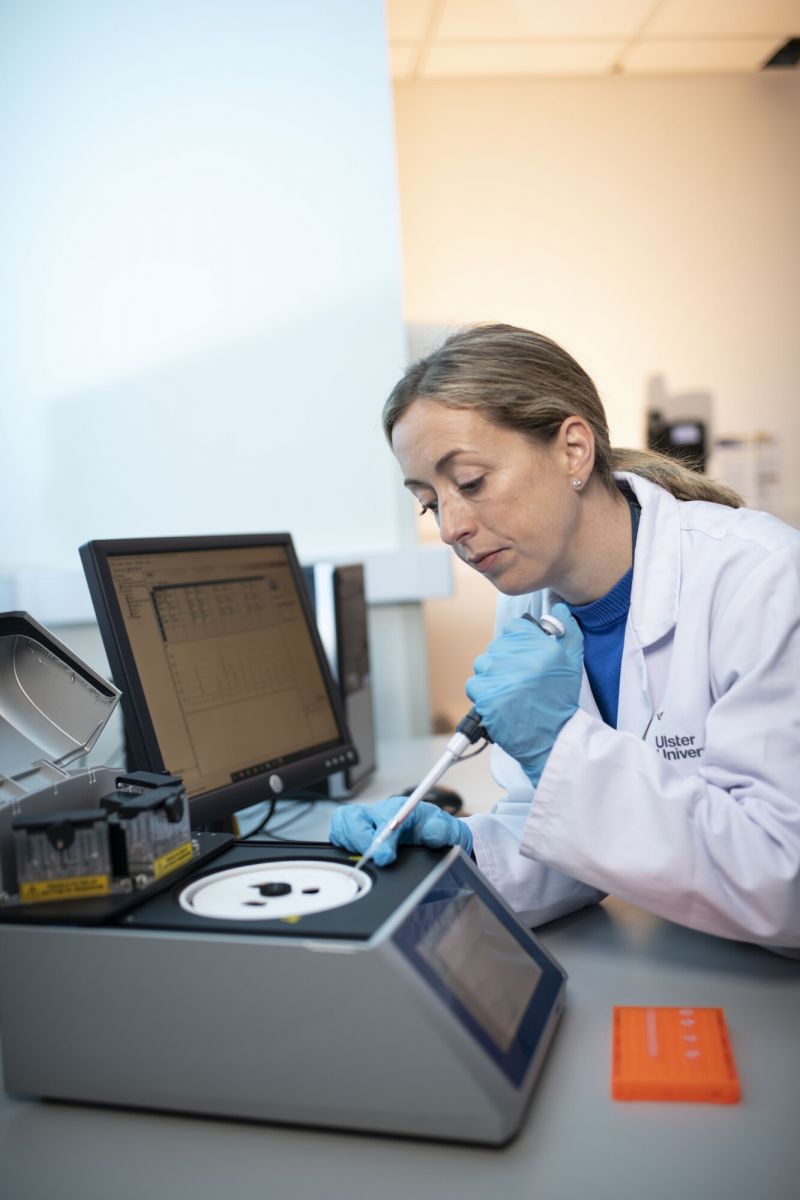

Cardiometabolic Epigenomics
Our research team use bioinformatic approaches to meta-analyse data with that from interdisciplinary projects at Ulster University....
Prostate Cancer
Investigating different pathways that contribute to prostate cancer development, with the aim of improving the diagnosis, prognosis and potential therapeutic intervention of this disease.
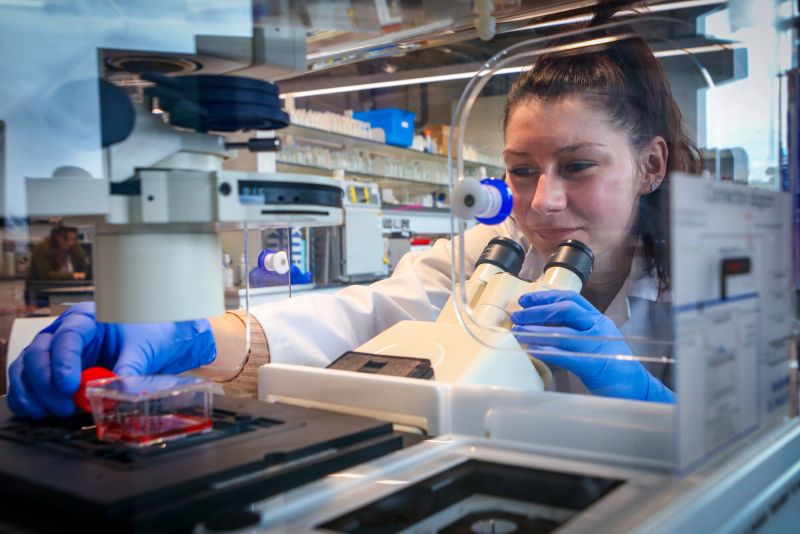

Prostate Cancer: Mechanisms of Disease
Investigating different pathways that contribute to prostate cancer development.
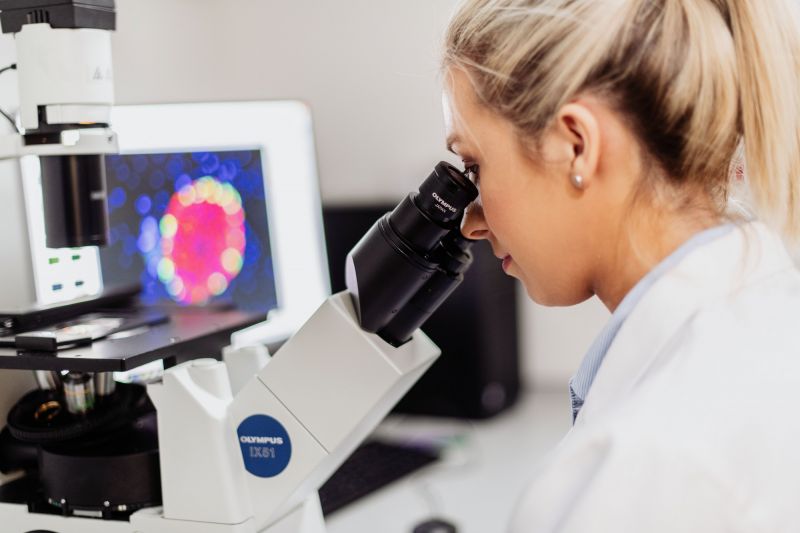
Digital histopathology for clinically applicable biomarker discovery
Aiming to discover novel ways to help clinicians make treatment decisions for prostate cancer patients using AI on cost effective...
Degenerative Eye Disease
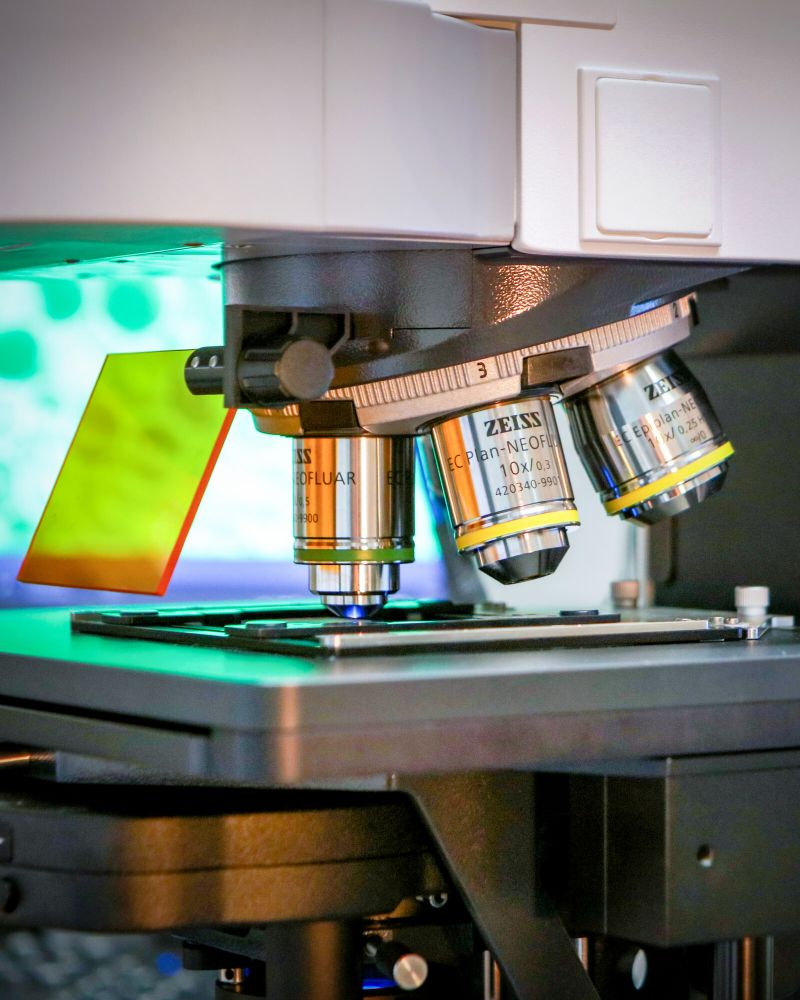

Glaucoma
Glaucoma is a progressive optic neuropathy characterised by the neurodegeneration of the retinal ganglion cells.
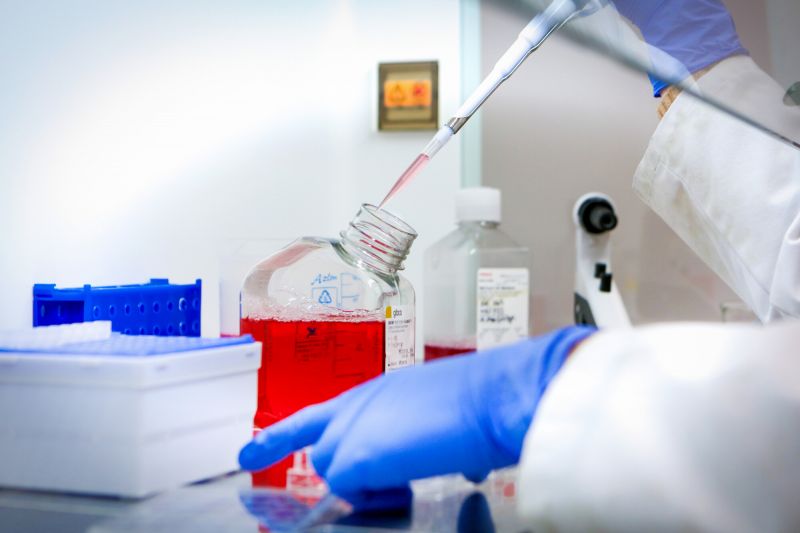
Corneal Dystrophy
Identifying the mutations which cause ocular surface disorders and in doing so they facilitate improved diagnosis and treatment in the...
Formulation & Manufacture of Personalised Medicines

Formulation and Manufacture of Personalised Medicines
This research has resulted in the development of an implantable drug delivery device (ChemoSeed®) for the local delivery of personalised...
















